Advanced examples¶
How to use Qibocal as a library¶
Qibocal also allows executing protocols without the standard interface.
In the following tutorial we show how to run a single protocol using Qibocal as a library. For this particular example we will focus on the t1_signal protocol (see also T1 experiments). The fastest way consists in using the Executor class in the following way
from qibocal.auto.execute import Executor
from qibocal.auto.mode import ExecutionMode
with Executor.open(
"myexec", # arbitrary name for executor
path="test_t1_signal", # path where the data will be stored
platform="my_platform", # platform to be used
targets=[0], # qubits on which the experiment will be executed
) as e:
# your experiments go here
The executor is responsible of running the routines on a platform and eventually store the history of multiple experiments. The context manager with provides an easy way to connect and disconnect from the platform.
In order to run an experiment the user needs to specify its parameters.
The user can check which parameters need to be provided either by checking the
documentation of the specific protocol or by simply inspecting protocol.parameters_type.
To run a t1_signal experiment is necessary to use invoke the protocol inside the with statement
output = e.t1_signal(delay_before_readout_start=0,
delay_before_readout_end=20_000,
delay_before_readout_step=50)
By default acquisition and fitting are performed.
The user can now use the raw data acquired by the quantum processor to perform an arbitrary post-processing analysis. This is one of the main advantages of this API compared to the cli execution.
Both the raw data and the fit data can be accessed from the history attribute of the Executor.
history = e.history
t1_res = history["t1_signal"][0]
data = t1_res.data # raw data
results = t1_res.results # fit data
In particular, the history object returns a dictionary that links the id of the experiments with the qibocal.auto.task.Completed object
How to add a new protocol¶
In this tutorial we show how to add a new protocol to Qibocal.
Protocol implementation in Qibocal¶
Currently, characterization/calibration protocols are divided in three steps: acquisition, fit and plot. Qibocal provides three data structures input parameters, data acquired and
results, that collect all the information concerning the routine.
The relationship between steps and data structures are summarized in the following bullets:
acquisitionreceives as inputparametersand outputsdatafitreceives as inputdataand outputsresultsplotreceives as inputdataandresultsto visualize the protocol
This approach is flexible enough to allow the data acquisition without performing a post-processing analysis.
Step by step tutorial¶
All protocols are located in qibocal.protocols.
Suppose that we want to code a protocol to perform a RX rotation for different
angles.
We create a file rotate.py in src/qibocal/protocols.
Parameters¶
First, we define the input parameters.
from dataclasses import dataclass
from ...auto.operation import Parameters
@dataclass
class RotationParameters(Parameters):
"""Parameters for rotation protocol."""
theta_start: float
"""Initial angle."""
theta_end: float
"""Final angle."""
theta_step: float
"""Angle step."""
nshots: int
"""Number of shots."""
In this case you define a range for the angle to be probed alongside the number of shots.
Note
It is advised to use dataclasses. If you are not familiar
have a look at the official documentation.
Data structure¶
Secondly, we define a data structure that aims at storing both the angles and the probabilities measured for each qubit. A generic data structure is usually composed of some raw data (the data attribute), which is usually coded as a dictionary of arrays plus additional information if required.
import numpy as np
import numpy.typing as npt
from dataclasses import dataclass, field
from ...auto.operation import Data
RotationType = np.dtype([("theta", np.float64), ("prob", np.float64)])
@dataclass
class RotationData(Data):
"""Rotation data."""
data: dict[QubitId, npt.NDArray[RotationType]] = field(default_factory=dict)
"""Raw data acquired."""
def register_qubit(self, qubit, theta, prob):
"""Store output for single qubit."""
ar = np.empty((1,), dtype=RotationType)
ar["theta"] = theta
ar["prob"] = prob
if qubit in self.data:
self.data[qubit] = np.rec.array(np.concatenate((self.data[qubit], ar)))
else:
self.data[qubit] = np.rec.array(ar)
Note
When the protocols will be executed the data will be saved automatically. The data attribute will be stored as a npz file, while the rest of the information will be stored as json file. If the user would like to use a custom format the implementation of a save method inside the data structure will be necessary.
Acquisition function¶
In the acquisition function we are going to perform the experiment.
Note
A generic acquisition function must have the following signature
from qibolab import Platform
from qibocal.auto.operation import QubitId, QubitPairId
from typing import Union
def acquisition(params: RoutineParameters, platform: Platform, targets: Union[list[QubitId], list[QubitPairId], list[list[QubitId]]]) -> RoutineData
"""A generic acquisition function."""
from qibolab import Platform
from qibocal.auto.operation import QubitId
def acquisition(
params: RotationParameters,
platform: Platform,
targets: list[QubitId],
) -> RotationData:
r"""
Data acquisition for rotation routine.
Args:
params (:class:`RotationParameters`): input parameters
platform (:class:`Platform`): Qibolab's platform
targets (list): list with target qubits
Returns:
data (:class:`RotationData`)
"""
# costruct range from RotationParameters
angles = np.arange(params.theta_start, params.theta_end, params.theta_step)
# create data structure
data = RotationData()
# create and execute circuit for each angle
for angle in angles:
circuit = Circuit(platform.nqubits)
for qubit in qubits:
circuit.add(gates.RX(qubit, theta=angle))
circuit.add(gates.M(qubit))
result = circuit(nshots=params.nshots)
for qubit in qubits:
# extract probability of 0
prob = result.probabilities(qubits=[qubit])[0]
# store measurements in Rotation Data
data.register_qubit(qubit, theta=angle, prob=prob)
return data
Result class¶
Here we decided to code a generic Results that contains the fitted parameters for each qubit.
from qibocal.auto.operation import QubitId
@dataclass
class RotationResults(Results):
"""Results object for data"""
fitted_parameters: dict[QubitId, list] = field(default_factory=dict)
Note
To check whether fitted parameters for a specific Qubit it might
be necessary to re-write the __contains__ method if the Results
inheritance include non-dictionary attributes.
Fit function¶
The following function performs a sinusoidal fit for each qubit.
Note
A generic fit function must have the following signature
def fit(data: RoutineData) -> RoutineResults """ A generic fit."
where Qubits is a dict[QubitId, Qubit].
from scipy.optmize import curve_fit
def fit(data: RotationData) -> RotationResults:
qubits = data.qubits
freqs = {}
fitted_parameters = {}
def cos_fit(x, offset, amplitude, omega):
return offset + amplitude * np.cos(omega*x)
for qubit in qubits:
qubit_data = data[qubit]
thetas = qubit_data.theta
probs = qubit_data.prob
popt, _ = curve_fit(cos_fit, thetas, probs)
freqs[qubit] = popt[2] / 2*np.pi
fitted_parameters[qubit]=popt.tolist()
return RotationResults(
fitted_parameters=fitted_parameters,
)
Report function¶
The report function generates a list of figures and an optional table to be shown in the html report. For the plotting function the user must use plotly in order to properly generate the report.
Note
A generic report function must have the following signature
import plotly.graph_objects as go
def plot(data: RoutineData, fit: RoutineResults, target: QubitId) -> list[go.Figure(), str]
""" A generic plotting function."""
The str in output can be used to create a table, which has 3 columns target, Fitting Parameter
and Value. Here is the syntax necessary to insert a raw in the table.
report = ""
target = 0
angle = 3.14
report += f" {qubit} | rotation angle: {angle:.3f}<br>"
This table can be omitted by returnig None.
Here is the plotting function for the protocol that we are coding:
import plotly.graph_objects as go
from qibocal.auto.operation import QubitId
def plot(data: RotationData, fit: RotationResults, target: QubitId):
"""Plotting function for rotation."""
figures = []
fig = go.Figure()
fitting_report = ""
qubit_data = data[target]
fig.add_trace(
go.Scatter(
x=qubit_data.theta,
y=qubit_data.prob,
opacity=1,
name="Probability",
showlegend=True,
legendgroup="Voltage",
),
)
if fit is not None:
fig.add_trace(
go.Scatter(
x=qubit_data.theta,
y=cos_fit(
qubit_data.theta,
*fit.fitted_parameters[target],
),
name="Fit",
line=go.scatter.Line(dash="dot"),
),
)
# last part
fig.update_layout(
showlegend=True,
xaxis_title="Theta [rad]",
yaxis_title="Probability",
)
figures.append(fig)
return figures, fitting_report
Create Routine object¶
rotation = Routine(acquisition, fit, plot)
"""Rotation Routine object."""
Add routine to Operation Enum¶
The last step is to add the routine that we just created to the available protocols in src/qibocal/protocols/__init__.py:
# other imports...
from rotate import rotation
__all__ = [
# other protocols....
"rotation",
]
Write a runcard¶
To launch the protocol a possible runcard could be the following one:
platform: my_platform
targets: [0,1]
actions:
- id: rotate
operation: rotation
parameters:
theta_start: 0
theta_end: 7
theta_step: 20
nshots: 1024
For more information about how to execute runcards see How to execute calibration protocols in Qibocal?.
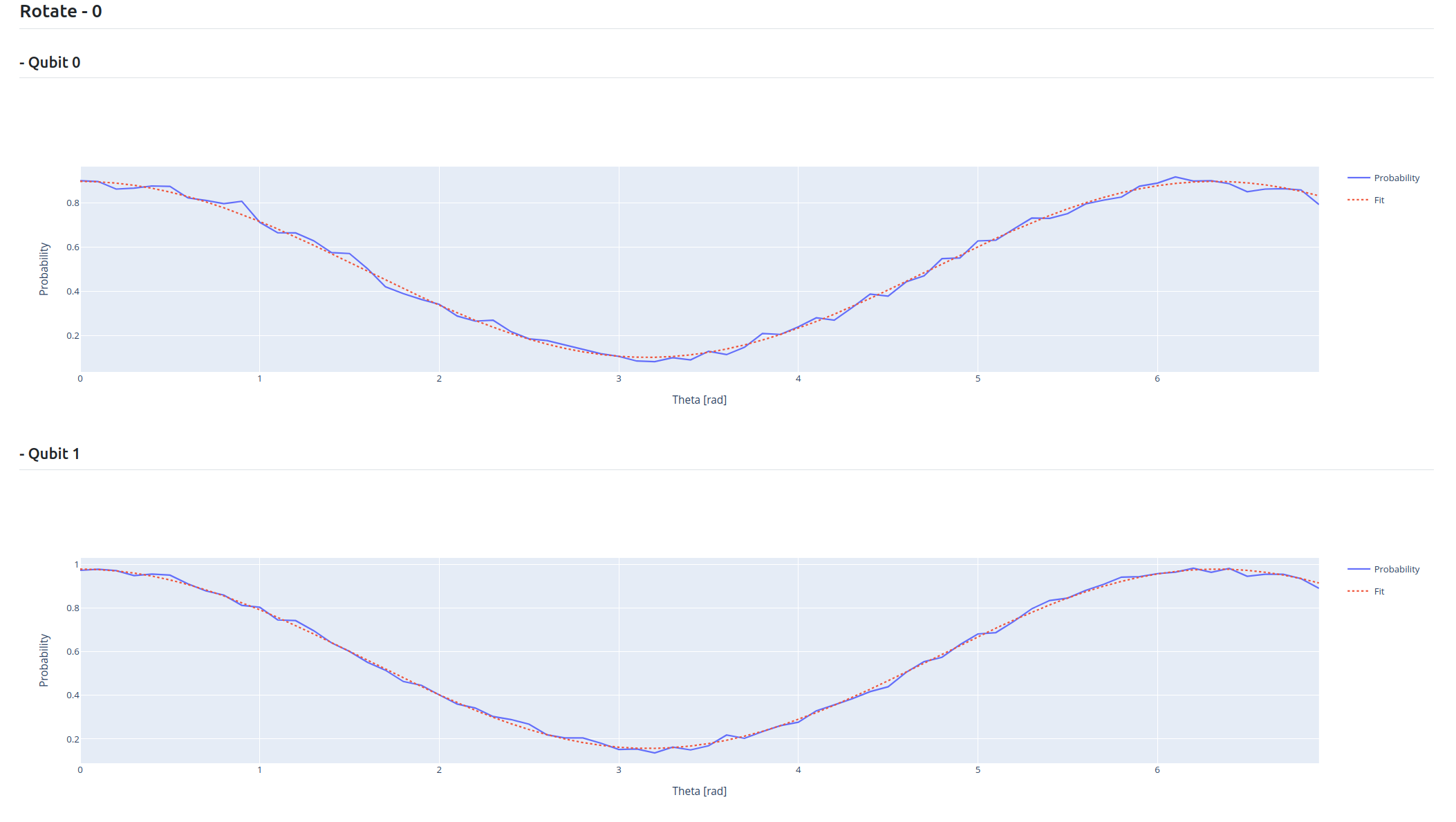
Here is the expected output: